Epigenetic research is entering a stage where chromatin profiling is increasingly defined by high-resolution, low-input technologies rather than traditional workflows.
Understanding gene regulation requires mapping DNA-protein interactions, identifying transcription factor binding sites, and profiling histone modifications. In many diseases—especially cancer—biological outcomes are not driven solely by genetic mutations. Epigenetic regulation plays a critical role.
Epigenomic technologies are now widely used across fields such as cancer research, stem cell and developmental biology, immunology, and single-cell genomics.
For more than two decades, researchers have relied on Chromatin Immunoprecipitation sequencing (ChIP-Seq) to study chromatin regulation. However, with the rise of low-input samples, single-cell technologies, and high-resolution epigenomics, traditional ChIP-Seq faces increasing challenges in experimental efficiency, sample requirements, and signal-to-noise ratio.
A new and powerful technique—Cleavage Under Targets and Tagmentation (CUT&Tag)—has rapidly gained attention as a more sensitive and efficient approach for chromatin profiling.
How did CUT&Tag rise so quickly? We’ll walk through the technological evolution from ChIP-Seq to CUT&Tag.
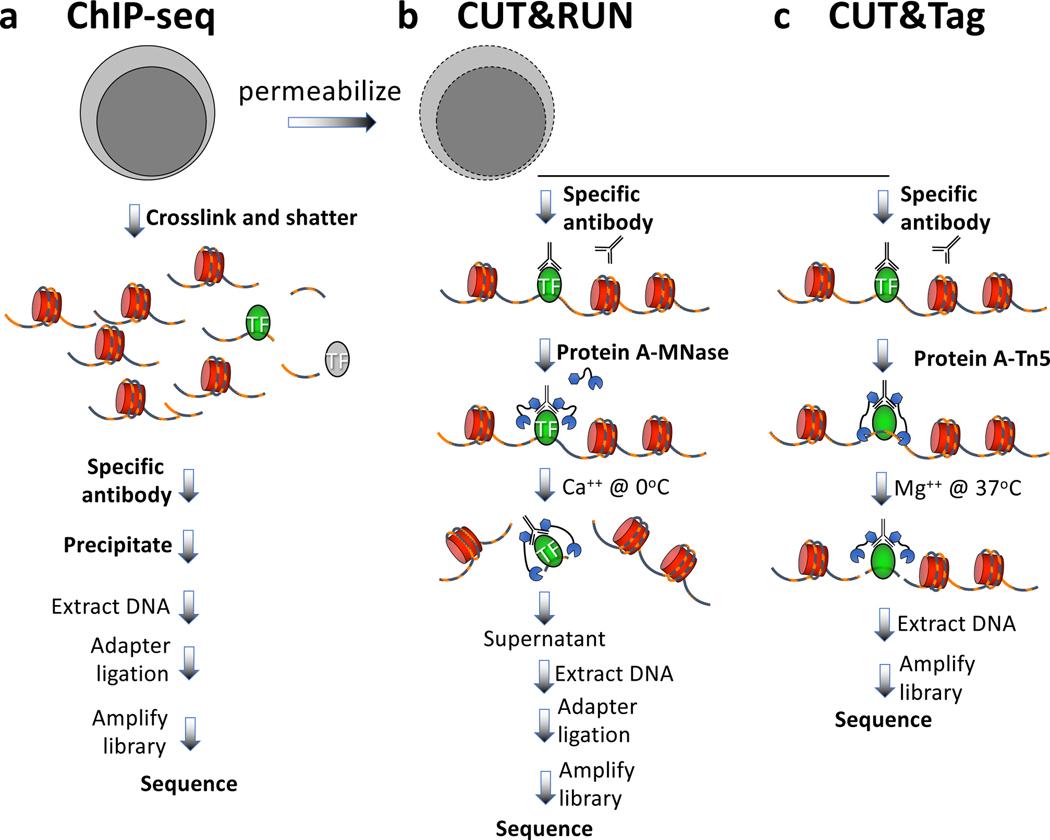
Fig.1 From ChIP-Seq to CUT&Tag: The evolution of chromatin profiling[1].
a. ChIP-Seq—the classic method: Genome-wide mapping of protein-DNA interactions; requires crosslinking, chromatin fragmentation, and relatively high cell input.
b. CUT&RUN—lower background: In situ cleavage using antibody-guided MNase, reducing background and improving signal specificity.
c. CUT&Tag—higher sensitivity: Targeted fragmentation with pAG-Tn5 enables high-sensitivity chromatin profiling with low input and a simplified workflow.
ChIP-Seq: The classic approach
ChIP-Seq enriches DNA fragments bound by specific proteins through immunoprecipitation, followed by high-throughput sequencing to identify binding sites across the genome. A typical workflow includes crosslinking, chromatin fragmentation, antibody immunoprecipitation, DNA purification, and library preparation[2].
Despite its widespread use, ChIP-Seq has several limitations:
• Long experimental time (typically 4 – 7 days)
• High sample input requirements
• Crosslinking variability that may introduce false positives or negatives
• Challenging optimization of chromatin fragmentation
• High background noise in sequencing data
As epigenetic research moves toward lower input, higher resolution, and more streamlined workflows, new technologies such as CUT&Tag have emerged to address these challenges.
CUT&Tag: A next-generation chromatin profiling method
CUT&Tag uses a Protein A/G-Tn5 transposase to bind antibodies and directly tag and cleave DNA at target sites in situ. After PCR amplification, sequencing-ready libraries can be generated directly[3].
Compared with ChIP-Seq, CUT&Tag offers several advantages:
• No crosslinking or sonication required
• Much lower background noise
• Higher signal-to-noise ratio
• Compatible with very low cell input (hundreds to thousands of cells, even ~10 cells)
• Fast workflow — completed within one day
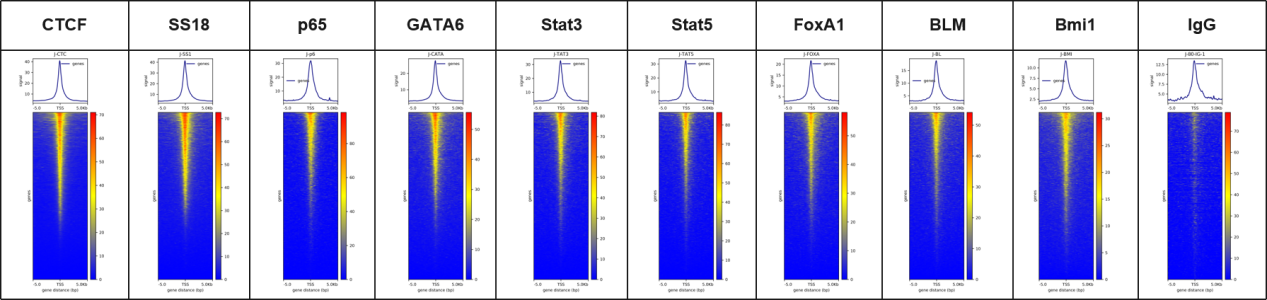
Fig. 2 CUT&Tag reveals cleaner chromatin profiles with a markedly higher signal-to-noise ratio.
With ultra-low input requirements, efficient library preparation, and superior sequencing efficiency, CUT&Tag is particularly well suited for rare samples and multi-omic studies, enabling high-resolution exploration of epigenetic regulation.
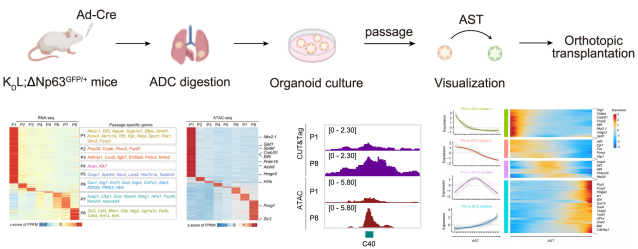
Fig. 3 Multi-omic profiling reveals resistance to KRAS inhibition in LKB1-mutant lung cancer.
As research questions become more complex, traditional ChIP-Seq often struggles to deliver optimal data quality. CUT&Tag, with its low input requirement, high sensitivity, and streamlined workflow, is rapidly emerging as a powerful new approach for epigenetic profiling.
If you are studying transcription factor binding, histone modifications, or epigenomics with limited input material, CUT&Tag offers an efficient and reliable solution.
1. Kaya-Okur, H. S., Janssens, D. H., Henikoff, J. G., Ahmad, K. & Henikoff, S. Efficient low-cost chromatin profiling with CUT&Tag. Nature Protocols 15, 3264–3283 (2020).
2. Johnson, D. S., Mortazavi, A., Myers, R. M. & Wold, B. Genome-wide mapping of in vivo protein–DNA interactions. Science 316, 1497–1502 (2007).
3. Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nature Communications 10, 1930 (2019).
4. Li, X. et al. Adeno-to-squamous transition drives resistance to KRAS inhibition in LKB1-mutant lung cancer. Nature 605, 764–773 (2022).
Media Contact
Company Name: Vazyme
Email: Send Email
Country: China
Website: https://www.vazymebiotech.com/